Abstract
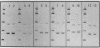
To facilitate genus and species level identification of a broad range of bacteria without the requirement of presumptive identification, we have developed a unified set of primers and polymerase chain reaction conditions to amplify spacer regions between the 16S and 23S genes in the prokaryotic rRNA genetic loci. Spacer regions within these loci show a significant level of length and sequence polymorphism across both genus and species lines. A generic pair of priming sequences was selected for the amplification of these polymorphisms from highly conserved sequences in the 16S and 23S genes occurring adjacent to these polymorphic regions. This single set of primers and reaction conditions was used for the amplification of 16S-23S spacer regions for over 300 strains of bacteria belonging to eight genera and 28 species or serotypes, including Listeria, Staphylococcus, and Salmonella species and additional species related to these pathogenic organisms. When the spacer amplification products were resolved by electrophoresis, the resulting patterns could be used to distinguish all of the species of bacteria within the test group. Unique elements in the amplification product patterns generally clustered at the species level, although some genus-specific characteristics were also observed. On the basis of the results obtained with our test group of 300 bacterial strains, amplification of the 16S-23S ribosomal spacer region is a suitable process for generating a data base for use in a polymerase chain reaction-based identification method, which can be comprehensively applied to the bacterial kingdom.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barry T., Colleran G., Glennon M., Dunican L. K., Gannon F. The 16s/23s ribosomal spacer region as a target for DNA probes to identify eubacteria. PCR Methods Appl. 1991 Aug;1(1):51–56. doi: 10.1101/gr.1.1.51. [DOI] [PubMed] [Google Scholar]
- Barry T., Powell R., Gannon F. A general method to generate DNA probes for microorganisms. Biotechnology (N Y) 1990 Mar;8(3):233–236. doi: 10.1038/nbt0390-233. [DOI] [PubMed] [Google Scholar]
- Bram R. J., Young R. A., Steitz J. A. The ribonuclease III site flanking 23S sequences in the 30S ribosomal precursor RNA of E. coli. Cell. 1980 Feb;19(2):393–401. doi: 10.1016/0092-8674(80)90513-9. [DOI] [PubMed] [Google Scholar]
- Brosius J., Dull T. J., Sleeter D. D., Noller H. F. Gene organization and primary structure of a ribosomal RNA operon from Escherichia coli. J Mol Biol. 1981 May 15;148(2):107–127. doi: 10.1016/0022-2836(81)90508-8. [DOI] [PubMed] [Google Scholar]
- Cedergren R., Gray M. W., Abel Y., Sankoff D. The evolutionary relationships among known life forms. J Mol Evol. 1988 Dec;28(1-2):98–112. doi: 10.1007/BF02143501. [DOI] [PubMed] [Google Scholar]
- Dams E., Hendriks L., Van de Peer Y., Neefs J. M., Smits G., Vandenbempt I., De Wachter R. Compilation of small ribosomal subunit RNA sequences. Nucleic Acids Res. 1988;16 (Suppl):r87–173. doi: 10.1093/nar/16.suppl.r87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox G. E., Stackebrandt E., Hespell R. B., Gibson J., Maniloff J., Dyer T. A., Wolfe R. S., Balch W. E., Tanner R. S., Magrum L. J. The phylogeny of prokaryotes. Science. 1980 Jul 25;209(4455):457–463. doi: 10.1126/science.6771870. [DOI] [PubMed] [Google Scholar]
- Garnier T., Canard B., Cole S. T. Cloning, mapping, and molecular characterization of the rRNA operons of Clostridium perfringens. J Bacteriol. 1991 Sep;173(17):5431–5438. doi: 10.1128/jb.173.17.5431-5438.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grimont F., Grimont P. A. Ribosomal ribonucleic acid gene restriction patterns as potential taxonomic tools. Ann Inst Pasteur Microbiol. 1986 Sep-Oct;137B(2):165–175. doi: 10.1016/s0769-2609(86)80105-3. [DOI] [PubMed] [Google Scholar]
- Göbel U. B., Geiser A., Stanbridge E. J. Oligonucleotide probes complementary to variable regions of ribosomal RNA discriminate between Mycoplasma species. J Gen Microbiol. 1987 Jul;133(7):1969–1974. doi: 10.1099/00221287-133-7-1969. [DOI] [PubMed] [Google Scholar]
- Kenerley M. E., Morgan E. A., Post L., Lindahl L., Nomura M. Characterization of hybrid plasmids carrying individual ribosomal ribonucleic acid transcription units of Escherichia coli. J Bacteriol. 1977 Dec;132(3):931–949. doi: 10.1128/jb.132.3.931-949.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiss A., Sain B., Venetianer P. The number of rRNA genes in Escherichia coli. FEBS Lett. 1977 Jul 1;79(1):77–79. doi: 10.1016/0014-5793(77)80354-2. [DOI] [PubMed] [Google Scholar]
- Lane D. J., Pace B., Olsen G. J., Stahl D. A., Sogin M. L., Pace N. R. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci U S A. 1985 Oct;82(20):6955–6959. doi: 10.1073/pnas.82.20.6955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehner A. F., Harvey S., Hill C. W. Mapping and spacer identification of rRNA operons of Salmonella typhimurium. J Bacteriol. 1984 Nov;160(2):682–686. doi: 10.1128/jb.160.2.682-686.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loughney K., Lund E., Dahlberg J. E. tRNA genes are found between 16S and 23S rRNA genes in Bacillus subtilis. Nucleic Acids Res. 1982 Mar 11;10(5):1607–1624. doi: 10.1093/nar/10.5.1607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Razin S., Gross M., Wormser M., Pollack Y., Glaser G. Detection of mycoplasmas infecting cell cultures by DNA hybridization. In Vitro. 1984 May;20(5):404–408. doi: 10.1007/BF02619586. [DOI] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Thomson-Carter F. M., Carter P. E., Pennington T. H. Differentiation of staphylococcal species and strains by ribosomal RNA gene restriction patterns. J Gen Microbiol. 1989 Jul;135(7):2093–2097. doi: 10.1099/00221287-135-7-2093. [DOI] [PubMed] [Google Scholar]
- Vilgalys R., Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol. 1990 Aug;172(8):4238–4246. doi: 10.1128/jb.172.8.4238-4246.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Fox G. E. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc Natl Acad Sci U S A. 1977 Nov;74(11):5088–5090. doi: 10.1073/pnas.74.11.5088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young R. A., Macklis R., Steitz J. A. Sequence of the 16 S-23 s spacer region in two ribosomal RNA operons of Escherichia coli. J Biol Chem. 1979 May 10;254(9):3264–3271. [PubMed] [Google Scholar]