Abstract
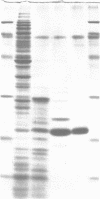
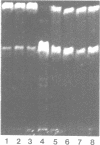
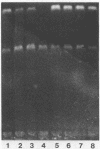
We have purified 3-methyladenine DNA glycosylase I from Escherichia coli to apparent physical homogeneity. The enzyme preparation produced a single band of Mr 22,500 upon sodium dodecyl sulphate/polyacrylamide gel electrophoresis in good agreement with the molecular weight deduced from the nucleotide sequence of the tag gene (Steinum, A.-L. and Seeberg, E. (1986) Nucl. Acids Res. 14, 3763-3772). HPLC confirmed that the only detectable alkylation product released from (3H)dimethyl sulphate treated DNA was 3-methyladenine. The DNA glycosylase activity showed a broad pH optimum between 6 and 8.5, and no activity below pH 5 and above pH 10. MgSO4, CaCl2 and MnCl2 stimulated enzyme activity, whereas ZnSO4 and FeCl3 inhibited the enzyme at 2 mM concentration. The enzyme was stimulated by caffeine, adenine and 3-methylguanine, and inhibited by p-hydroxymercuribenzoate, N-ethylmaleimide and 3-methyladenine. The enzyme showed no detectable endonuclease activity on native, depurinated or alkylated plasmid DNA. However, apurinic sites were introduced in alkylated DNA as judged from the strand breaks formed by mixtures of the tag enzyme and the bacteriophage T4 denV enzyme which has apurinic/apyrimidinic endonuclease activity. It was calculated that wild-type E. coli contains approximately 200 molecules per cell of 3-methyladenine DNA glycosylase I.
Full text
PDF













Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Behmoaras T., Toulmé J. J., Hélène C. A tryptophan-containing peptide recognizes and cleaves DNA at apurinic sites. Nature. 1981 Aug 27;292(5826):858–859. doi: 10.1038/292858a0. [DOI] [PubMed] [Google Scholar]
- Beranek D. T., Weis C. C., Swenson D. H. A comprehensive quantitative analysis of methylated and ethylated DNA using high pressure liquid chromatography. Carcinogenesis. 1980 Jul;1(7):595–606. doi: 10.1093/carcin/1.7.595. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Clarke N. D., Kvaal M., Seeberg E. Cloning of Escherichia coli genes encoding 3-methyladenine DNA glycosylases I and II. Mol Gen Genet. 1984;197(3):368–372. doi: 10.1007/BF00329931. [DOI] [PubMed] [Google Scholar]
- Demple B., Linn S. DNA N-glycosylases and UV repair. Nature. 1980 Sep 18;287(5779):203–208. doi: 10.1038/287203a0. [DOI] [PubMed] [Google Scholar]
- Evensen G., Seeberg E. Adaptation to alkylation resistance involves the induction of a DNA glycosylase. Nature. 1982 Apr 22;296(5859):773–775. doi: 10.1038/296773a0. [DOI] [PubMed] [Google Scholar]
- Kaasen I., Evensen G., Seeberg E. Amplified expression of the tag+ and alkA+ genes in Escherichia coli: identification of gene products and effects on alkylation resistance. J Bacteriol. 1986 Nov;168(2):642–647. doi: 10.1128/jb.168.2.642-647.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karran P., Hjelmgren T., Lindahl T. Induction of a DNA glycosylase for N-methylated purines is part of the adaptive response to alkylating agents. Nature. 1982 Apr 22;296(5859):770–773. doi: 10.1038/296770a0. [DOI] [PubMed] [Google Scholar]
- Karran P., Lindahl T., Ofsteng I., Evensen G. B., Seeberg E. Escherichia coli mutants deficient in 3-methyladenine-DNA glycosylase. J Mol Biol. 1980 Jun 15;140(1):101–127. doi: 10.1016/0022-2836(80)90358-7. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Laval J. Two enzymes are required from strand incision in repair of alkylated DNA. Nature. 1977 Oct 27;269(5631):829–832. doi: 10.1038/269829a0. [DOI] [PubMed] [Google Scholar]
- Lindahl T. DNA repair enzymes. Annu Rev Biochem. 1982;51:61–87. doi: 10.1146/annurev.bi.51.070182.000425. [DOI] [PubMed] [Google Scholar]
- Lindahl T. New class of enzymes acting on damaged DNA. Nature. 1976 Jan 1;259(5538):64–66. doi: 10.1038/259064a0. [DOI] [PubMed] [Google Scholar]
- Lindahl T., Nyberg B. Rate of depurination of native deoxyribonucleic acid. Biochemistry. 1972 Sep 12;11(19):3610–3618. doi: 10.1021/bi00769a018. [DOI] [PubMed] [Google Scholar]
- Nakabeppu Y., Kondo H., Sekiguchi M. Cloning and characterization of the alkA gene of Escherichia coli that encodes 3-methyladenine DNA glycosylase II. J Biol Chem. 1984 Nov 25;259(22):13723–13729. [PubMed] [Google Scholar]
- Nakabeppu Y., Miyata T., Kondo H., Iwanaga S., Sekiguchi M. Structure and expression of the alkA gene of Escherichia coli involved in adaptive response to alkylating agents. J Biol Chem. 1984 Nov 25;259(22):13730–13736. [PubMed] [Google Scholar]
- Nakabeppu Y., Yamashita K., Sekiguchi M. Purification and characterization of normal and mutant forms of T4 endonuclease V. J Biol Chem. 1982 Mar 10;257(5):2556–2562. [PubMed] [Google Scholar]
- Nakatsu Y., Nakabeppu Y., Sekiguchi M. Action of T4 endonuclease V on polydeoxyribonucleotides with apyrimidinic or apurinic sites. J Biochem. 1982 Jun;91(6):2057–2065. doi: 10.1093/oxfordjournals.jbchem.a133899. [DOI] [PubMed] [Google Scholar]
- Pierre J., Laval J. Specific nicking of DNA at apurinic sites by peptides containing aromatic residues. J Biol Chem. 1981 Oct 25;256(20):10217–10220. [PubMed] [Google Scholar]
- Riazuddin S., Lindahl T. Properties of 3-methyladenine-DNA glycosylase from Escherichia coli. Biochemistry. 1978 May 30;17(11):2110–2118. doi: 10.1021/bi00604a014. [DOI] [PubMed] [Google Scholar]
- Sakumi K., Nakabeppu Y., Yamamoto Y., Kawabata S., Iwanaga S., Sekiguchi M. Purification and structure of 3-methyladenine-DNA glycosylase I of Escherichia coli. J Biol Chem. 1986 Nov 25;261(33):15761–15766. [PubMed] [Google Scholar]
- Sedgwick B. Molecular cloning of a gene which regulates the adaptive response to alkylating agents in Escherichia coli. Mol Gen Genet. 1983;191(3):466–472. doi: 10.1007/BF00425764. [DOI] [PubMed] [Google Scholar]
- Seeberg E., Nissen-Meyer J., Strike P. Incision of ultraviolet-irradiated DNA by extracts of E. coli requires three different gene products. Nature. 1976 Oct 7;263(5577):524–526. doi: 10.1038/263524a0. [DOI] [PubMed] [Google Scholar]
- Seeberg E. Reconstitution of an Escherichia coli repair endonuclease activity from the separated uvrA+ and uvrB+/uvrC+ gene products. Proc Natl Acad Sci U S A. 1978 Jun;75(6):2569–2573. doi: 10.1073/pnas.75.6.2569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinum A. L., Seeberg E. Nucleotide sequence of the tag gene from Escherichia coli. Nucleic Acids Res. 1986 May 12;14(9):3763–3772. doi: 10.1093/nar/14.9.3763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas L., Yang C. H., Goldthwait D. A. Two DNA glycosylases in Escherichia coli which release primarily 3-methyladenine. Biochemistry. 1982 Mar 16;21(6):1162–1169. doi: 10.1021/bi00535a009. [DOI] [PubMed] [Google Scholar]
- Valerie K., Henderson E. E., deRiel J. K. Identification, physical map location and sequence of the denV gene from bacteriophage T4. Nucleic Acids Res. 1984 Nov 12;12(21):8085–8096. doi: 10.1093/nar/12.21.8085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto Y., Katsuki M., Sekiguchi M., Otsuji N. Escherichia coli gene that controls sensitivity to alkylating agents. J Bacteriol. 1978 Jul;135(1):144–152. doi: 10.1128/jb.135.1.144-152.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]